Help on the information displayed in GeneNest
How to access GeneNest databases?
You can go to the clusters directly via the GeneNest on
http://GeneNest.molgen.mpg.de
page using the "Quick Search". The following searches can be performed:
- Clustername
- Clustername.Contigname
- Clone ID
- GenBank Accession Number
Use an empty query string to go to the first cluster in a database. For more search options use the "Complex Search Form"
Detailed searches
On the Complex Search Form you have several options to search against a GeneNest database. You can search with:
- Clustername or Clustername.Contigname
- Clone ID
- GenBank accession number
- Annotation in gene descriptions
- Library ID
- Use a DNA or protein sequence in fasta or plain text format to blast against consensus database.
Except for annotation search or blast you can seperate several queries by comma and the results are combined using OR.
The annotation field may use Regular Expressions using the special symbols `$', `^', ` * ', `[', `]', `|',
`(', `)', `!', and `\'.
Visualization of a cluster
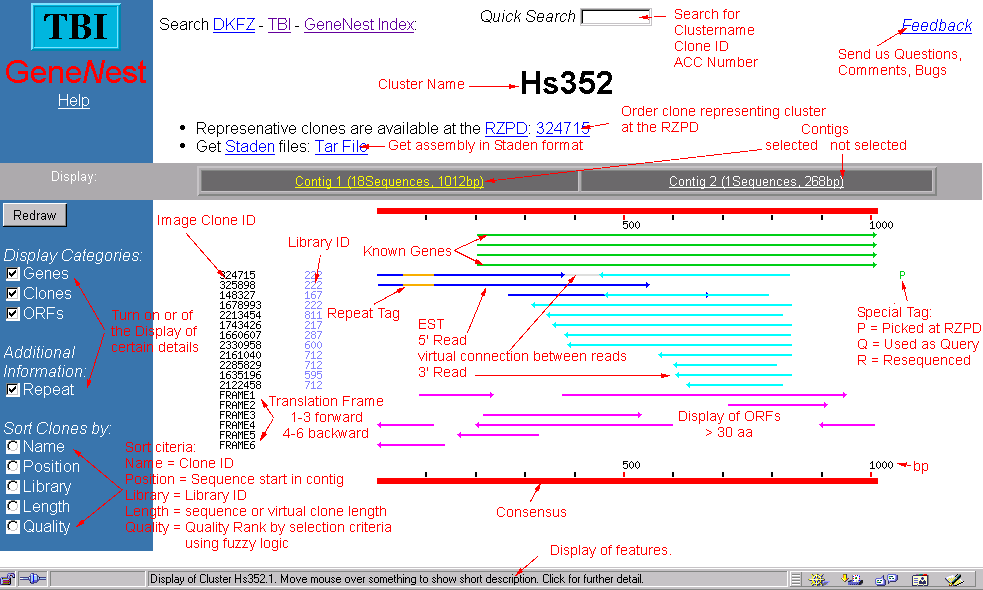
The information for a cluster as well as some further analysis are displayed in a graphical overview. a cluster of
highly similar sequences might fall apart into different contigs during the procedure of assembly (see
General help on EST clustering ).
Usually you can obtain some more information about the element by moving with the mouse over the element. By clicking to an
element you usually link to a new page with information about the object.
View Example
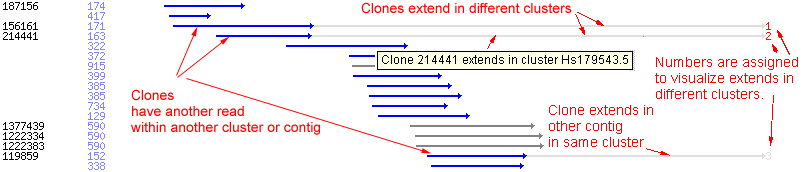
Clone Extends
A cDNA clone might have several EST sequences. if these sequences are not linked together by sequence overlap they might fall
into different clusters or contigs of the same cluster. ESTs within the same cluster belonging to the same clone are connected
in the graphical display and a virtual sequence of the complete clone is computed. Clones which have ESTs in other clusters or
contigs are extended towards the end of the consensus and a tag is put at the end which links to the other contig or cluster.
Extends in other clusters can mean that these clusters should belong to the same Gene but the sequence information of the
ESTs is to short to detect overlap. Or it might be a sign that the clone sequences are concatenated and belong to different
genes.
Further questions - Feedback
Tim Beissbarth
- Last Change: 7.03.00